Pantothenate kinase–associated neurodegeneration is an autosomal recessive disorder characterized by accumulation of iron mainly in the basal ganglia.1,2 In about half of these cases, patients have an identifiable mutation in the PANK2 gene.1
We previously described a 13-year-old boy who showed the “eye of the tiger” sign on a T2-weighted magnetic resonance (MR) image3 that is highly specific not only for this disease but also for a mutation in the PANK2 gene.1 Here we report on our screening for mutations of the PANK2 gene conducted on the genomic DNA of the patient and his family (Fig. 1).
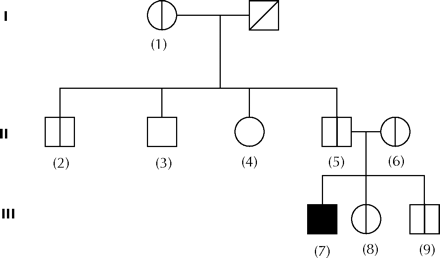
Fig. 1: Pedigree of our patient with pantothenate kinase–associated neurodegeneration. Circles represent females and squares, males. The square with an oblique line through it represents a deceased grandparent. The patient (homozygous for the mutation H173Y in the PANK2 gene) is represented by a black box. Heterozygotes for the mutation are shown with half-cut boxes. Homozygotes for the wild-type genotype are shown with blank boxes. Numerical codes for all subjects are given in parentheses.
DNA was isolated from peripheral blood using a phenol-chloroform reference protocol. All exons of the gene were amplified by polymerase chain reaction (PCR),4 and the amplified products were subjected to cycle sequencing using an ABI PRISM 377 (Applied Biosystems) DNA sequencer. A novel mutation was identified in exon 2 resulting in a change of histidine to tyrosine at amino acid position 173 (H173Y), which also creates an RsaI restriction site. To better visualize the restriction products, exon 2 of the gene was re-amplified using custom-designed primers (5'-ACCTGACCTCCAAT GTGG-3') and (5'-AGTGTGGA GACTCGAGAAG-3'). Amplified products (176 base pairs) were subjected to restriction endonuclease analysis by using RsaI; 0.5 units of enzyme were used for 10 mL of PCR reaction products. After a 3-hour incubation at 37°C, electrophoresis was carried out on a 2% agarose gel. Fig. 2 shows the restriction analysis for all the family members. The patient is homozygous for this mutation, whereas both the siblings and subjects 1, 2, 5 and 6 are heterozygous.
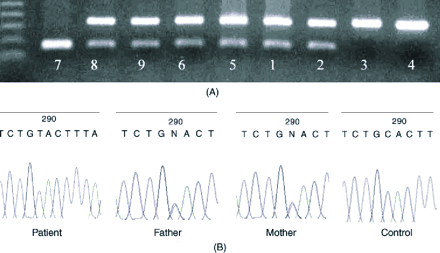
Fig. 2: The mutation H173Y (C to T) creates an RsaI restriction site. (A) A 2% agarose gel shows restriction analysis by the RsaI enzyme. The amplified product (176 base pairs) of exon 2 generated by custom- designed primers was digested by the enzyme at 37°C for 3 hours. The numbers below each lane represent numerical codes of subjects in Fig. 1. Subjects 3 and 4 carry the wild-type genotype and, thus, their amplified products remained undigested, showing only 1 band in the top panel. Both the siblings (8 and 9) and subjects 1, 2, 5 and 6 are heterozygous for this mutation, with 1 allele carrying the mutation and the other carrying the wild-type gene; thus, the top band shows the wild-type allele and the bottom one shows the mutant allele. Patient (7) is homozygous for the mutation, with both the alleles having the mutation, thus, displaying only 1 lower band. (B) Electrophoretogram of patient, father, mother and control. A transition from “C” in control to “T” in the patient can be seen, whereas the 2 parents are heterozygous.